Like the one ring of power in Tolkien's "Lord of the Rings," deoxyribonucleic acid (DNA) is the master molecule of every cell. It contains vital information that gets passed on to each successive generation. It coordinates the making of itself as well as other molecules (proteins). If it is changed slightly, serious consequences may result. If it is destroyed beyond repair, the cell dies.
Changes in the DNA of cells in multicellular organisms produce variations in the characteristics of a species. Over long periods of time, natural selection acts on these variations to evolve or change the species.
Advertisement
The presence or absence of DNA evidence at a crime scene could mean the difference between a guilty verdict and an acquittal. DNA is so important that the United States government has spent enormous amounts of money to sequence DNA in the human genome in hopes of understanding and finding cures for many genetic diseases. Finally, from the DNA of one cell, we can clone an animal, a plant or perhaps even a human being.
But what is DNA? Where is it found? What makes it so special? How does it work? In this article, we will look deep into the structure of DNA and explain how it makes itself and how it determines all of your traits. First, let's look at how DNA was discovered.
DNA is one of a class of molecules called nucleic acids. Nucleic acids were originally discovered in 1868 by Friedrich Miescher, a Swiss biologist, who isolated DNA from pus cells on bandages. Although Miescher suspected that nucleic acids might contain genetic information, he could not confirm it.
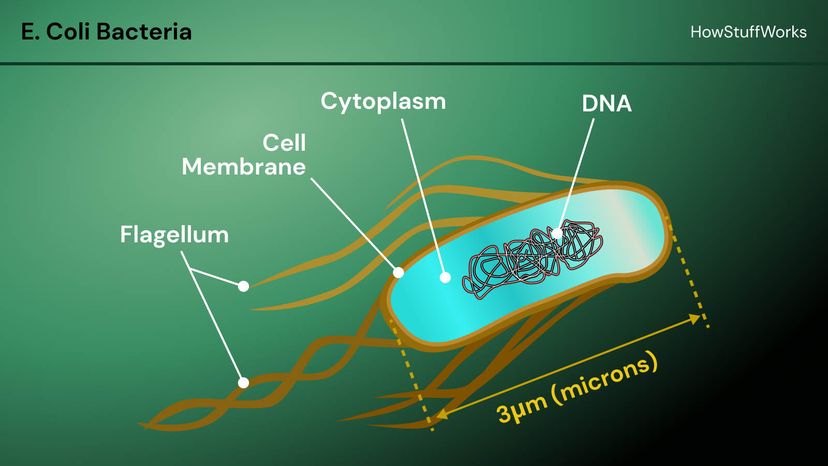
In 1943, Oswald Avery and colleagues at Rockefeller University showed that DNA taken from a bacterium, Streptococcus pneumonia, could make noninfectious bacteria become infectious. These results indicated that DNA was the information-containing molecule in the cell. The information role of DNA was further supported in 1952 when Alfred Hershey and Martha Chase demonstrated that to make new viruses, a bacteriophage virus injected DNA, not protein, into the host cell.
So, scientists had theorized about the informational role of DNA for a long time, but nobody knew how this information was encoded and transmitted. Many scientists guessed that the structure of the molecule was important to this process. In 1953, James D. Watson and Francis Crick discovered the structure of DNA at Cambridge University.
Basically, Watson and Crick used molecular modeling techniques and data from other investigators (including Maurice Wilkins, Rosalind Franklin, Erwin Chargaff and Linus Pauling) to solve the structure of DNA. Watson, Crick and Wilkins received the Nobel Prize in Medicine in 1962 for the discovery (Franklin, who was Wilkins' collaborator and provided a key piece of data that revealed the structure to Watson and Crick, died before the prize was awarded).
Advertisement